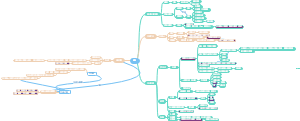
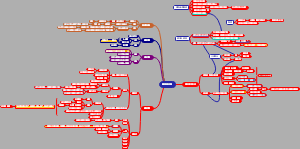
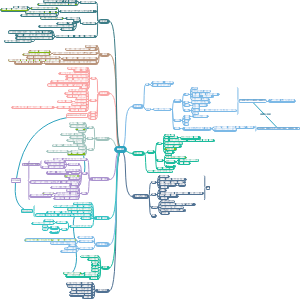
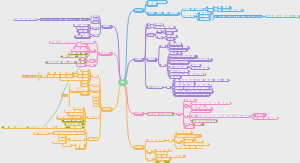
导图社区 合成生物学进展时间线
- 423
- 16
- 2
- 举报
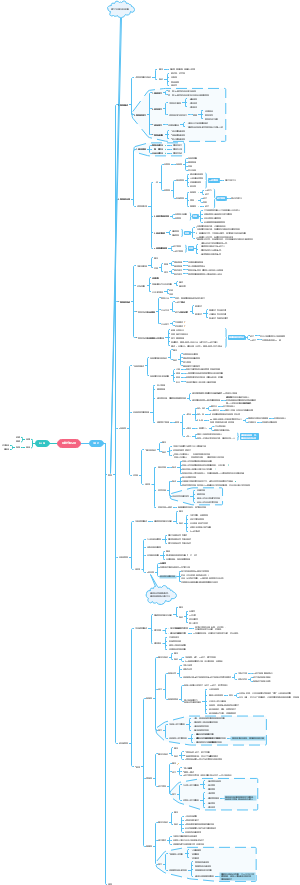
合成生物学进展时间线
超全!最细节的合成生物学进展时间线梳理图,梳理了从1960s乳糖操纵子的研究到2020年蛋白质结构预测与设计的过程和时间点。
编辑于2021-11-30 11:41:55- 合成生物学
- 相似推荐
- 大纲
合成生物学
1960s
乳糖操纵子的研究
1970s
分子克隆技术发展
1980s
"组学"的发展
1990s
DNA测序技术的发展
2000
第一个合成开关—生物开关和振荡子
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000). Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
自动调节的负反馈线路
Becskei, A. & Serrano, L. Engineering stability in gene networks by autoregulation. Nature 405, 590–593 (2000)
2001
基于群体效应的细胞间的通信线路
Weiss, R. & Knight, T. F. Jr. in DNA Computing (eds Condon, A. & Rozenberg, G.) 1–16 (Springer, 2001).
拓展大肠杆菌的遗传密码
Wang, L., A. Brock, B. Herberich, and P. G. Schultz. “Expanding the Genetic Code of Escherichia Coli.” Science 292, no. 5516 (April 20, 2001): 498–500. https://doi.org/10.1126/science.1060077.
2002
最早实现遗传网络的组合合成
Guet, C. C., Elowitz, M. B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1466–1470 (2002).
发展用于转录研究的合成线路
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73(2002). Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002). Blake, W. J., Kaern, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003).
2003
大肠杆菌中实现青蒿素前体途径的工程化
Martin, V. J., Pitera, D. J., Withers, S. T., Newman, J. D. & Keasling, J. D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nature Biotech. 21, 796–802 (2003).
2004
SB 1.0:第一次合成生物学国际会议
麻省理工学院举行第一届iGEM竞赛
RNA装置用于基因表达的调节模块
Isaacs, F. J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nature Biotech. 22, 841–847 (2004).
2005
大肠杆菌中的光传感电路设计
Levskaya, A. et al. Synthetic biology: engineering Escherichia coli to see light. Nature 438, 441–442 (2005).
由RNA调控的可编程的基因表达
Bayer, T. S. & Smolke, C. D. Programmable ligand controlled riboregulators of eukaryotic gene expression. Nature Biotech. 23, 337–343 (2005).
生成多细胞模式的线路
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
2006
利用工程菌侵入癌细胞
Anderson, J. C., Clarke, E. J., Arkin, A. P. & Voigt, C. A. Environmentally controlled invasion of cancer cells by engineered bacteria. J. Mol. Biol. 355, 619–627 (2006).
DNA折纸
Rothemund P W K. Folding DNA to create nanoscale shapes and patterns[J]. Nature, 2006, 440(7082): 297-302.
2007
改造噬菌体水解细菌的生物膜
Lu, T. K. & Collins, J. J. Dispersing biofilms with engineered enzymatic bacteriophage. Proc. Natl Acad. Sci. USA 104, 11197–11202 (2007).
2008
利用大肠杆菌中氨基酸的代谢生产生物燃料
Atsumi, S., Hanai, T. & Liao, J. C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89 (2008).
可用于逻辑运算的RNA装置
Win, M. N. & Smolke, C. D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008).
快速、可调节的合成基因振荡器
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
2009
Gibson等对DNA组装的研究
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nature Methods 6, 343–345 (2009)
MAGE的研究
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009)
设计可计算的线路
Friedland, A. E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
边缘检测的合成与线路的工程化
Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009).
2010
可编辑控制微生物致死的开关
Callura, J. M., Dwyer, D. J., Isaacs, F. J., Cantor, C. R. & Collins, J. J. Tracking, tuning, and terminating microbial physiology using synthetic riboregulators. Proc. Natl Acad. Sci. USA 107, 15898–15903 (2010).
人工合成细菌染色体
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
利用基因时钟同步细胞群的耦合振动波
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
2011
在大肠杆菌中建立一套完整的布尔逻辑门
Tamsir, A., Tabor, J. J. & Voigt, C. A. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011).
人工合成酵母的部分基因组
Dymond, J. S. et al. Synthetic chromosome arms function in yeast and generate phenotypic diversity by design. Nature 477, 471–476 (2011).
2012
设计出复杂的多层基因环路
Moon, T. S., Lou, C., Tamsir, A., Stanton, B. C. & Voigt, C. A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
动态代谢流控制生物柴油的生产
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor–regulator system for production of chemicals and fuels derived from fatty acids. Nature Biotech. 30, 354–359 (2012).
DNA数据储存
Church G M, Gao Y, Kosuri S. Next-generation digital information storage in DNA[J]. Science, 2012, 337(6102): 1628-1628.
CRISPR/Cas9体外特征鉴定
Gasiunas G, Barrangou R, Horvath P, Siksnys V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A 2012; 109:E2579-2586. Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012; 337:816-821
2013
酵母商业化生产青蒿素
Paddon, C. J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
CRISPR用于人细胞编辑
Mali P, Yang L, Esvelt K M, et al. RNA-guided human genome engineering via Cas9[J]. Science, 2013, 339(6121): 823-826. Cong L, Ran F A, Cox D, et al. Multiplex genome engineering using CRISPR/Cas systems[J]. Science, 2013, 339(6121): 819-823.
CRISPR用于调控
Gilbert L A, Larson M H, Morsut L, et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes[J]. Cell, 2013, 154(2): 442-451.
2014
扩展生命遗传密码(碱基)
Malyshev D A, Dhami K, Lavergne T, et al. A semi-synthetic organism with an expanded genetic alphabet[J]. Nature, 2014, 509(7500): 385-388.
合成酵母染色体计划
Gibson D G, Venter J C. Synthetic biology: Construction of a yeast chromosome[J]. Nature, 2014, 509(7499): 168-169.
基于纸的基因线路
Pardee K, Green A A, Ferrante T, et al. based synthetic gene networks[J]. Cell, 2014, 159(4): 940-954.
CRISPR用于筛选
Wang T, Wei J J, Sabatini D M, et al. Genetic screens in human cells using the CRISPR-Cas9 system[J]. Science, 2014, 343(6166): 80-84. Shalem O, Sanjana N E, Hartenian E, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells[J]. Science, 2014, 343(6166): 84-87.
2015
设计酵母合成吗啡
Galanie S, Thodey K, Trenchard I J, et al. Complete biosynthesis of opioids in yeast[J]. Science, 2015, 349(6252): 1095-1100.
微生物群遗传振荡
Chen Y, Kim J K, Hirning A J, et al. Emergent genetic oscillations in a synthetic microbial consortium[J]. Science, 2015, 349(6251): 986-989.
2016
人工合成最小细菌
Hutchison C A, Chuang R Y, Noskov V N, et al. Design and synthesis of a minimal bacterial genome[J]. Science, 2016, 351(6280).
计算设计人工蛋白
Huang P S, Boyken S E, Baker D. The coming of age of de novo protein design[J]. Nature, 2016, 537(7620): 320-327.
合成57密码子大肠杆菌
Ostrov N, Landon M, Guell M, et al. Design, synthesis, and testing toward a 57-codon genome[J]. Science, 2016, 353(6301): 819-822.
人造胰岛β细胞
Xie M, Ye H, Wang H, Charpin-El Hamri G, Lormeau C, Saxena P, Stelling J, Fussenegger M: b-Cell-mimetic designer cells provide closed-loop glycemic control. Science 2016, 354:1296-1301.
2017
制造出“稳定”的半合成有机体
Zhang Y, Ptacin J L, Fischer E C, et al. A semi-synthetic organism that stores and retrieves increased genetic information[J]. Nature, 2017, 551(7682): 644-647.
DNA储存数码信息新方法
Erlich Y, Zielinski D. DNA Fountain enables a robust and efficient storage architecture[J]. Science, 2017, 355(6328): 950-954. Shipman S L, Nivala J, Macklis J D, et al. CRISPR–Cas encoding of a digital movie into the genomes of a population of living bacteria[J]. Nature, 2017, 547(7663): 345-349.
重新设计合成了5条酵母染色体
Richardson S M, Mitchell L A, Stracquadanio G, et al. Design of a synthetic yeast genome[J]. Science, 2017, 355(6329): 1040-1044.
CRISPR用于记录
Sheth R U, Yim S S, Wu F L, et al. Multiplex recording of cellular events over time on CRISPR biological tape[J]. Science, 2017, 358(6369): 1457-1461. Schmidt F, Cherepkova M Y, Platt R J. Transcriptional recording by CRISPR spacer acquisition from RNA[J]. Nature, 2018, 562(7727): 380-385.
CRISPR用于检测
Gootenberg J S, Abudayyeh O O, Lee J W, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2[J]. Science, 2017, 356(6336): 438-442.
2018
人工合成首例单染色体酵母菌株
Shao Y, Lu N, Wu Z, et al. Creating a functional single-chromosome yeast[J]. Nature, 2018, 560(7718): 331-335.
Biobits合成生物学教育产品
Huang A, Nguyen P Q, Stark J C, et al. BioBits™ Explorer: A modular synthetic biology education kit[J]. Science advances, 2018, 4(8): eaat5105. Stark J C, Huang A, Nguyen P Q, et al. BioBits™ Bright: A fluorescent synthetic biology education kit[J]. Science advances, 2018, 4(8): eaat5107.
人工构建多细胞自组装结构
Toda, Satoshi, Lucas R. Blauch, Sindy K. Y. Tang, Leonardo Morsut, and Wendell A. Lim. “Programming Self-Organizing Multicellular Structures with Synthetic Cell-Cell Signaling.” Science, May 31, 2018, eaat0271. https://doi.org/10.1126/science.aat0271. Glass, David S., and Ingmar H. Riedel-Kruse. “A Synthetic Bacterial Cell-Cell Adhesion Toolbox for Programming Multicellular Morphologies and Patterns.” Cell 174, no. 3 (July 2018): 649-658.e16. https://doi.org/10.1016/j.cell.2018.06.041.
酶法DNA合成的突破
Palluk S, Arlow D H, De Rond T, et al. De novo DNA synthesis using polymerase-nucleotide conjugates[J]. Nature biotechnology, 2018, 36(7): 645.
2019
定向进化能利用CO2的大肠杆菌
Gleizer S, Ben-Nissan R, Bar-On Y M, et al. Conversion of Escherichia coli to generate all biomass carbon from CO2[J]. Cell, 2019, 179(6): 1255-1263. e12.
八碱基生命体
Hoshika S, Leal N A, Kim M J, et al. Hachimoji DNA and RNA: A genetic system with eight building blocks[J]. Science, 2019, 363(6429): 884-887.
从头设计蛋白质药物
Silva D A, Yu S, Ulge U Y, et al. De novo design of potent and selective mimics of IL-2 and IL-15[J]. Nature, 2019, 565(7738): 186-191.
从头设计蛋白开关
Langan R A, Boyken S E, Ng A H, et al. De novo design of bioactive protein switches[J]. Nature, 2019, 572(7768): 205-210. Ng A H, Nguyen T H, Gómez-Schiavon M, et al. Modular and tunable biological feedback control using a de novo protein switch[J]. Nature, 2019, 572(7768): 265-269.
设计酵母合成大麻
Luo X, Reiter M A, d’Espaux L, et al. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast[J]. Nature, 2019, 567(7746): 123.
2020
蛋白质结构预测与设计
Chen Z, Kibler R D, Hunt A, et al. De novo design of protein logic gates[J]. Science, 2020, 368(6486): 78-84.
自动化设计与实验
Erlich Y, Zielinski D. DNA Fountain enables a robust and efficient storage architecture[J]. Science, 2017, 355(6328): 950-954. Shipman S L, Nivala J, Macklis J D, et al. CRISPR–Cas encoding of a digital movie into the genomes of a population of living bacteria[J]. Nature, 2017, 547(7663): 345-349.
绿色制造
Luo X, Reiter M A, d’Espaux L, et al. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast[J]. Nature, 2019, 567(7746): 123.
工程微生物医疗
Silva D A, Yu S, Ulge U Y, et al. De novo design of potent and selective mimics of IL-2 and IL-15[J]. Nature, 2019, 565(7738): 186-191.
2021
等待揭晓
Chen Z, Kibler R D, Hunt A, et al. De novo design of protein logic gates[J]. Science, 2020, 368(6486): 78-84.
文化
其他应用
生命拓展
医疗应用
技术
全基因组工程
代谢工程
线路工程