导图社区 Frontier Tech _ Lifescience _ Ab_Ph.D. Pipeline
- 31
- 0
- 0
- 举报
Frontier Tech _ Lifescience _ Ab_Ph.D. Pipeline
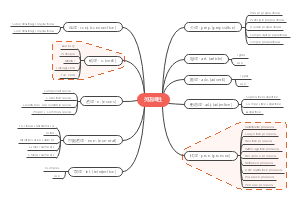
这是一篇关于Ph.D. Pipeline的思维导图,主要内容包括:reported library,Epitope mapping,background,VL library,Fab library,VHH library,Ab label,General Method,reagent,Ab graft,pioneering uses,scFv library。
编辑于2024-09-18 19:01:42- 酵母
- General Method
- 生物医药行业简介
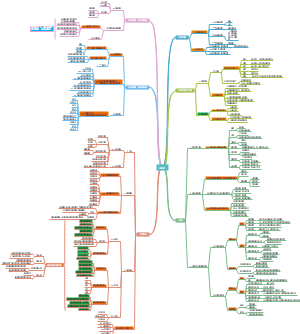
这是一篇关于生物医药行业简介的思维导图,从行业分布、发展趋势、就业方向等角度梳理。主要内容包括:延申,职业,科服,诊断,制药。
- Frontier Tech _ Lifescience _ Ab_Ph.D. Pipeline
这是一篇关于Ph.D. Pipeline的思维导图,主要内容包括:reported library,Epitope mapping,background,VL library,Fab library,VHH library,Ab label,General Method,reagent,Ab graft,pioneering uses,scFv library。
- Frontier Tech _ Ab RD
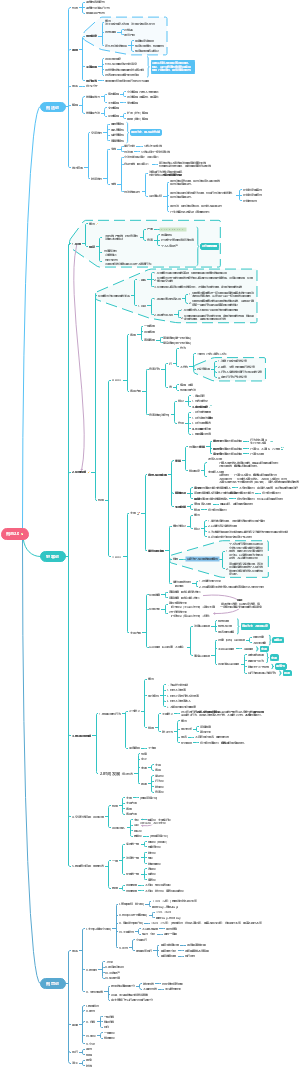
这是一篇关于Ab RD (details mean everythin的思维导图,主要内容包括:Structure,Sources,Quality Control,in vivo assay,Cell based functional assay,Phage based VH/VL screen,Method,Paper,Properity,Protein,Molecule。
Frontier Tech _ Lifescience _ Ab_Ph.D. Pipeline
社区模板帮助中心,点此进入>>
- 生物医药行业简介
这是一篇关于生物医药行业简介的思维导图,从行业分布、发展趋势、就业方向等角度梳理。主要内容包括:延申,职业,科服,诊断,制药。
- Frontier Tech _ Lifescience _ Ab_Ph.D. Pipeline
这是一篇关于Ph.D. Pipeline的思维导图,主要内容包括:reported library,Epitope mapping,background,VL library,Fab library,VHH library,Ab label,General Method,reagent,Ab graft,pioneering uses,scFv library。
- Frontier Tech _ Ab RD
这是一篇关于Ab RD (details mean everythin的思维导图,主要内容包括:Structure,Sources,Quality Control,in vivo assay,Cell based functional assay,Phage based VH/VL screen,Method,Paper,Properity,Protein,Molecule。
- 相似推荐
- 大纲
Ph.D. Pipeline
Yeast display 与 Phage display 相比,酵母表面展示具有以下显著优势: 1、酵母具有类似于高等真核生物的分泌途径, 蛋白质折叠在内质网中,其中伴侣蛋白,折叠酶和质量控制机制确保仅分泌正确折叠的蛋白质。而噬菌体是原核系统,有些抗体对大肠杆菌有毒的克隆,其生长缓慢甚至不生长,导致文库的偏向与克隆的丢失。 2、通过噬菌体展示筛选更高亲和力的抗体时,通常会受到筛选处理过程的不利影响,不仅取决于亲和力,还取决于抗体表达水平。而酵母展示系统采用FACS分选技术,基于抗体亲和力及展示水平进行筛选,因此消除了表达引起的偏差,且能区分相差很小亲和力的克隆。 3、酵母展示是多价展示,能够一次通过Sorting直接分选高,中&低亲和力的克隆,非常方便 4、通过使用双染色的 FACS,可以直接在酵母细胞表面上确定抗体亲和力,从而无需耗时的亚克隆、表达和纯化。 5、抗体筛选速度快,3min即可完成10万个细胞的分选,获得100+的unique sequence 候选克隆。 6、在相同的展示系统下可对抗体的亲和力和稳定性进行优化。 from: 成都阿帕克生物科技有限公司
General Method
Phmd design
Phagemid vector 1, for Human Name > Author > Reference pSEX81, Welschof-1997. pHAL, pHAL1, M. Hust-2005. parameters affecting the display of Ab on phage. pHAL5, pHAL6, 2, for mouse 3, for Shark library HEL-5A7:type1 vNAR produced by Helen Dooley from an immunized library against lysozyme. paper= 2003-selection & characterization of Naturally occuring IgNAR from immunized Sharks by Ph.D.
subclone
V-region subclone into phagemid vector refer to paper=2020-Generation of a Large Peptide Phage Display Library by Self-Ligation of Whole-Plasmid PCR Product
position
N-peptide-short linker(GSG/GGG)-amber stop-p3 C-p3-short linker(GSG/GGG)-peptide
helper
identify features of hyperphage hypophage helper phage
production
Production after selection 1, format > vector > cells human IgG1, pCSEH,pCSL, HEK293. scFv-(mouse IgG2a)Fc, pCSE2.6, HEK293
quantify
Traditional method: Progeny phages are often visualized as plaques, or holes, in a lawn of bacteria on an agar-filled Petri dish. Newly method: enumerate phage by fluorescent signal.
diversity
bio-panning
Affinity
Principle Method of apparatus (BLI)Gatorbio —— gator (BLI)Fortebio —— octet (SPR)Cytiva —— Biacore 8K/8K+/S200/T200/X100 binding affinity determined by fluorescence polarization in heinis 2020 paper.
Biacore (SPR)
Gator (BLI)
ForteBio Ocet (BLI)
ForteBio is part of Sartorius group
子主题
peptide library
Library production refer to NEB Lib, how they evaluate Lib? phagemid design 、selection、optimization evaluation: 1) quality of library, 2) type of repertoire (eg: naive, synthetic, semisynthetic, ……) 3) diversity (NNK, NNS, KKK, ……) 4) performance of library (positive clone per panning, hit rate, affinity, developability) 5) effective size (codon optimization, open read frame, non-toxic) concept Alan Perelson [ 34 , 35 ] formalized this concept as p = e −Np , P is the probability that a given antibody does not recognize an epitope of a random shape, N is the size of the antibody library, p is the probability that said antibody contacts said epitope with certain affinity. If p = 5 µ M (5 × 10 −6 M), which is a weak dissociation constant, but measurable and differentiable from non-specific binding, a library of N = 10 6 antibodies would yield a p = 6.8 × 10 −3 ( ≈ 2.7 −(1,000,000 × 0.000005) ). This means that the universe of virtually all possible random epitopes would be recognized by an antibody with affinity around 5 µ M in a library of one million antibody variants. To reach the same (low) p value but with a dissociation constant of 5 nM (5 × 10 −9 M), N should be 10 9 antibody variants. Hence, the larger the library, the higher the chances that an antibody will specifically bind a random epitope with higher affinity. refer to : 2019(Review)-Phage Display Libraries for Antibody Therapeutic Discovery and Development.
phage library
Refer to NEB "M13KE" vector,it`s not necessary to express antibiotic genes before plating or to use helper, for random peptide library , pahge vector may better. design a better phage vector.
phagemid design
helper design
refer to superphage(phagenomics) & hyperphage(dakewe)
try other virus
scFv library
Andy Q. Yuan 2020-Isolation of and Characterization of Neutralizing Antibodies to Covid-19 from a Large Human Naïve scFv Phage Display Library. Jianhui Cai 2021-Construction and application of a human scFv phage display library based on Cre‑LoxP recombination for anti‑PCSK9 antibody selection. Dario Neri 2014-A Highly Functional Synthetic Phage Display Library Containing over 40 Billion Human Antibody Clones.
scFv
principle to design synthetic scFv library 1, determin the wanted V-region (could be 1 or 3 or 6 or …… of them) common order: CDR3 >> CDR2 > CDR1, VH >> VK > VL. 2, fix framework of wanted V-region eg: FW3, FW2, FW1 of VH, or * of VL 3, design virable form of library, thus got theoretical diversity, eg: (NNK)10 (would be flanked by homologous region or restrication enzyme site to circlize) 4, construct into phagemid, 5, electroporation into F+ E.coli strain
traditional method
whole-plasmid PCR & circlize
one step
primary + Cre-LoxP
Jianhui Cai 2021-Construction and application of a human scFv phage display library based on Cre‑LoxP recombination for anti‑PCSK9 antibody selection.
Framework determination
Virable region selection
VHH library
Ying Tianlei, 2020-Identification of Human Single-Domain Antibodies against SARS-CoV2. Dimiter S. Dimitrov, 2008-Human domain antibodies to conserved sterically restricted regions on gp120 as exceptionally potent cross-reactive HIV-1 neutralizers. 2020-Potent neutralization of SARS-CoV-2 by human antibody heavy-chain variable domains isolated from a large library with a new stable scaffold.
Fab library
2013-A fully synthetic human Fab antibody library based on fixed VH/VL frameworkpairin gs with favorable biophysical properties. 2012-General strategy for the generation of human antibody variable domains with increased aggregation resistance.
VL library
2018-Productive common light chain libraries yield diverse panels of high affinity bispecific antibodies.
pioneering uses
VHH on bacteria
VHH on red blood cells
VHH on phage
VHH based gene therapy
2020-Humanized single domain antibodies neutralize SARS-CoV-2 by targeting the spike receptor binding domain.pdf
reagent
reagent for 1. basic research 1.1 new binder discovery 1.2 detection reagent, eg: labled nanoBody 2. industrial detection 2.1 Biopharmed company source: news from (account=iDiscover, name=鲎试剂停产,内毒素检测难题来了) thought: 2.1.1 production of 鲎试剂 by E.coli/CHO/cell-free ? 2.1.2 detect endotoxin via NanoBody ? method can refer to paper=2020-Pyruvate dehydrogenase complex-enzyme2, a new target …… 2.2 Food industry 2.3 Agricultural idustry 3. clinical diagnostic 3.1 supplier 3.2 CRO
Ab graft
Ab graft (human & animal) 1, scFv 2, VHH 3, Fab 4, IgG1/4 5, IgE/M 6, vNAR 7, mutispecific 8, Fc 2017-IgG Fc domains that bind C1q but not effector Fcγ receptors delineate the importance of complement-mediated effector functions. 9, CH2 / CH3 / -Fc
Knob in Hole
ADC
Fc
Ab label
Ab Label 1, Biotiny-streptavdin-beads 1) protein Labeling with the IMPACT ™Kit, 【protein with a C-terminal thioester , Peptide with an N-terminal cysteine(peptide can contain a fluorescent label, modified amino acid, biotin, etc.) 】 General Protocol 1. One of the components should have a final concentration of at least 0.5–1 mM. For the ligation of a peptide to a protein we use 0.01–0.1 mM protein with 0.5–1 mM peptide. 2. Combine the two components in the presence of 0.1 M Tris, pH 8.5, 0.1–0.5 M NaCl and 10 mM MESNA and incubate overnight at 4°C. Alternatively, the reaction can be incubated at 25°C for 1–4 hours. 3. The ligation may be visualized by a 10% or 12% SDS-PAGE as a shift in mobility of the ligated protein, or can easily be detected by Western blot using an antibody specific for the peptide. Add 3X SDS Sample Buffer (with DTT) to the protein sample, boil for 5 minutes and analyze by SDS-PAGE, with unligated protein as a control (Figure 2). (refer to https://www.neb.com/products/n6707-ptxb1-vector#Protocols,%20Manuals%20&%20Usage) 2) 2013-Generating conformation-specific synthetic antibodies to trap proteins in selected functional states. 2, Fluorescent dye label 3, HRP paper:1)2020-Nanobody‑horseradish peroxidase and -EGFP fusions as reagents to detect porcine parvovirus in the immunoassays. 4,
background
Phage vs.
a brief summary in .xls file, file path as below: OneDrive >> 个人 >> 文档 >> 新建Microsoft Excel工作表 M13 λ T4 T7 f1 fd
Lib vs.
Phage display library comparison 1, library classfication 2, library feature 3, principle for choose
display method vs.
display method comparison 1, Ab display —— biomarker discovery & Ab generation 2, OFReome display —— target identification 3, single-gene display —— epitope characterization
Epitope mapping
Epitope Mapping == Identify binding motif 【structure & seq analysis】 eg:structure & sequence based analysis ligands/peptide/Ab/endogenous molecule/small molecules resources: 1) http://www.gpcrdb.org 2) https://zhanglab.dcmb.med.umich.edu/GPCR-EXP/ 2021-Functional GLP-1R antibodies identified from a synthetic GPCR-focused library demonstrate potent blood glucose control. 2015-Subunit disassembly and inhibition of TNFα by a semi-synthetic bicyclic peptide 2021-Versatile and multivalent nanobodies efficiently neutralize SARS-CoV-2 2021-Llamanade: an open-source computational pipeline for robust nanobody humanization. 2019-AppA: a web server for analysis, comparison, and visualization of contact residues and interfacial waters of antibody-antigen structures and models. conformational specific Ab 2013-Generating conformation-specific synthetic antibodies to trap proteins in selected functional states. 一、 Relationship between Protein sequence, structure and function. •https://www.ucl.ac.uk/biosciences/departments/structural-and-molecular-biology •http://www.bioinf.org.uk/research.html • Amino acid sequence alignment •ImMunoGeneTics(IMGT)numbering system is used. (2008-Human domain antibodies to conserved sterically restricted regions on gp120 as exceptionally potent cross-reactive HIV-1 neutralizers_SUP) • VectorDB contains annotations and sequence information for many vectors commonly used in molecular biology. • http://genome-www.stanford.edu/vectordb/vector.html 二、 •蛋白二级结构预测:http://www.compbio.dundee.ac.uk/jpred/ •跨膜区预测:http://www.cbs.dtu.dk/services/TMHMM-2.0/ •信号肽预测:http://www.cbs.dtu.dk/services/SignalP/ •糖基化位点预测:http://www.cbs.dtu.dk/services/NetOGlyc/ •理化性质分析:https://web.expasy.org/protparam/ •亲水性疏水性分析:https://web.expasy.org/cgi-bin/protscale/protscale.pl • 三、 •SWISSMODEL server (in the automated mode) •HADDOCK server (for Molecular docking) •PISA server (residues Analysis on the interface) •Developability Index (to Screen Ab Aggregation Propensity) •SAbPred (http://opig.stats.ox.ac.uk/webapps/newsabdab/sabpred/ ) •AppA web server for analysis, comparison, and visualization of contact residues and interfacial waters of antibody-antigen structures and models. •Llamanade (an open-source computational pipeline for robust nanobody humanization) •PyIgClassify (database and web server that provides access to assignments of all CDR structures in the PDB to our classification system. Python-based Ig classification available at http://dunbrack2.fccc.edu/pyigclassify/ ) 四、
reported library
Naive
2 naive library generated by Xoma: 1) XFab1 2) XscFv2
XFab1
XscFv2
Immune
Synthetic
Semisynthetic